Page 54 - 《水产学报》2025年第11期
P. 54
胡利,等 水产学报, 2025, 49(11): 119105
chit1 LOC105331572 Cht Cht Cht 基序符号
92 Glyco_18 BD2 BD2 BD2
0 100 200 300 400 500 600 700 motif symbol
83 chi3l1-2 LOC105328391 Glyco_18 Cht BD2
motif 1
0 100 200 300 400 motif 2
70 chit1l LOC105331571 Glyco_18 Cht BD2 motif 3
motif 4
0 100 200 300 400 500 motif 5
amcase-2 LOC105331570 LDLa Glyco_18 Cht BD2
29 motif 6
0 100 200 300 400 500 600 700 motif 7
pchi LOC105332575 Glyco_18 motif 8
0 100 200 300 400 motif 9
42 100 Glyco_18 motif 10
pchi1 LOC105332577 motif 11
0 100 200 300 400 motif 12
amcase-1 LOC105332622 Glyco_18 Cht BD2 Cht BD2
0 100 200 300 400 500 600 700
100 64 LOC10533891 Glyco_18
chi3l1-1
0 100 200 300 400
uncharacterized LOC10533257 Glyco_18 Glyco_18 Glyco_18 Glyco_18 Glyco_18 Cht BD2 Cht BD2
0 200 400 600 800 1 000 1 200 1 400 1 600 1 800 2 000 2 200
ctbs-1 LOC105347316 Glyco_18
0 100 200 300 400
ctbs-2 LOC105348542 Glyco_18
0 100 200 300
1.00
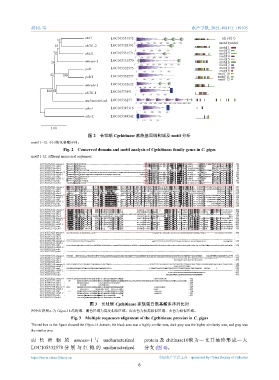
图 2 长牡蛎 Cgchitinase 家族基因结构域及 motif 分析
motif 1~12. 不同的氨基酸序列。
Fig. 2 Conserved domain and motif analysis of Cgchitinase family genes in C. gigas
motif 1-12. different amino acid sequences.
LOC105332622 amcase-1: MSRLNLPSLLVFLVILKVSH-----------------------------------------------------------SYMRVCYYTNWAQYRPNNG-KY-VPENLDPYLC-SHLIFAFAKM-NGNRL-----VAFEWNDESTDWMRGMYAKFNDIKLKNPTVKTLLAV : 102
LOC105331571 chit1l : MARLLFTFIYLTYLCHTAAL-----------------------------------------------------------KYKRVCYYTNWAQYRNGIA-KF-EPSHIDPSLC-THIIYAFGKL-EDNGI-----TNFEWNDET------RYVEVNAFKRNNTGLKTLLAM : 96
LOC105347316 ctbs-1 : MNITRIIFLTIHLGLGVLSE------------------------------------------------------------------YIVKDIAKE-------CPC--DPVYCRTHKYYYNREV-VVVAEGKSDQSAWHWHD--------ITMVITTEEVGKEEAKKLMCT : 86
LOC105348542 ctbs-2 : MGREFFCPVTIVFVFSLIVFIVN--------------------------------------------------------GTKTSCPCATEQQCEV-------IKDDK------RKEIFIFS-L-ENSKDKW---SRYDWSKVTTVVMVG-YINLELMCFAHKHGARAVLI : 95
LOC105331570 amcase-2: MKHLLRSSLIPVIFVLVRDLSLVLGQPRVFSFTPRRGTSCRPSEFHCGNGTCIPEEWACDRDSNCVDGKDESDSLCDTKKHRRVCYYTNWAQYREAPA-RF-TPEDINPDLC-THLIYAFAEL-RGAQI-----ETSEWNDPQ------MYERFNRIKNKNREIKTLLAV : 155
LOC105338914 chi3l1-1 : MKHNWTDRQTNCKVHYQVKIDQQLERPFSDLQSKMADILKRTVVQFILWTVVTFQ--------------------SEASGFKRVCYYTNWSESRAVVDFRFHLKEHLDPFLC-THLIYAFANI-NPTELRIE--RVYSSEDDGKVPGSGLMFDFTALKKKNPKLKTILSI : 146
LOC105331572 chit1 : -MAGFKHLVTTLLVCSLASIGLA--------------------------------------------------------EYIMVCYYTNWAQYRNGVG-KF-MPKDIDPNLC-THIIYAFGKL-DGNKI-----TNFEWNDNDDYIN--LYKQVNEHKAANPNLKTLLAM : 102
LOC105328391 chi3l1-2 : MMVNWTSISFLILLFFSSGYG------------------------------------------------------------MLVCYFTNWSQYRPGNG-RF-TPQNIDPHLC-THIAYAFGKLGSNNRV-----ATVEQNDVT------LYSQVNALKSTNPNLKTLLAM : 96
LOC105332575 pchi : MNVYTIVSVFVTVISLTFVQGQ---------------------------------------------------------HFQRFCVYNGFAIARDGEH-AL-QPEDIDPFLC-THIVFSFAEV-DETGTRLKDPDHFEANY--------LYQRIIRLRKLNHRLNMILSV : 101
LOC105332577 pchi1 : -MKGQACMVLISVWLSCVDG-----------------------------------------------------------EFIKMCYYSAWAIYRDGSQ-AL-TPEDIDAHHC-SHLVLAYATI-DDTGKNLKFPDTYENMH--------LPERLNDMRDHKDDLNMVLSV : 98
LOC105332622 amcase-1: GGWNMGSKPFTQMVKTPESRA-----EFTKSTIKFLRERNFDGLDLDWEYPAN--RGSPKEDKDRFTKLVIQLRSAFNQEAAMTGKPPLLLTAAVAAGKDKVDTGYDVATIAKHMDFINVMTYDLHGPWEK--VTGHNSPLHSRRDEMGPDTQLNIAWAAEYWYKLGLPK : 263
LOC105331571 chit1l : GGWTAGSKPYSDMAASPENRR-----TFINASISWLRKYDFDGLDMDWEYPAN--RGGVPEDFNNFPILLKEILEAFTEEAKTSGKSRLLLTAAVGVGKSVADTAYNIPEMSKYLDFISLMAYDLRGGWEK--TTGFNAALY--RSSADSSDEYNVAFAVDYWLRKGTPK : 255
LOC105347316 ctbs-1 : AHQNHANFAFIREVPTITNTSDPVLLDWLEETVKKEKELYADYVALDLLKL----LGQCKADSSHVQEVLALLK--FMVEKFTSPSAELVCIVPYKPPCYDGDCSLTQYMAEHCAAFITSPESFITCDTNPKCIAKATMPMT------------SLVFGVEEFIDHGITP : 238
LOC105348542 ctbs-2 : ANFPKAN------LTNATDRS-----KWVKKNLQTVQKNYLDGINFDFEDFIE---ENRTDQRDGYTSLVNETFQTFKQANK---NYQVTVDVAWSP-VGVDSRFYDYLKLSYVSDFLFIMSYDEQSQIYGECVAGANSGFY------------KTLSGFYKYYEMGADL : 235
LOC105331570 amcase-2: GGWNAGSGAFSSMVASTESRL-----LFVNSAIDFLRKYDFDGLDLDWEYPTR--RGGRPEDKENFAKLVKLLREAFDREARRTGRPVLLLSAAVPTSKELVDQAYDIITLGKYLDFINLMTYDFHGGWES--TVGHNSPLYSRRSETEAQRRMNTNWAARYWNQNGVPK : 316
LOC105338914 chi3l1-1 : GGEHSGE--FQLIMGSDKNIK-----TFAQNIYIYIRDRNFDGIDIDWEYPGA-------IYKNEFVIFLKELRAVFAAEAKQTGNEEKSISIAGAPGISNIETSYNITEIVKYVDFINVMAYDYSGPWSH--KTGFMAPLYSRSSDKSFDQTLSQNWTMNKYLELGAPP : 300
LOC105331572 chit1 : GGWTAGSLIYSNMASTAANRK-----EFIDSAIDWLRKYNFDGLDMDWEYPAN--RDGKPYDKENFAHLCKETREAFDNEAQTTGRDRLLFTSAVGAGKAVVDSAYDVPVMAQYMDFICIMSYDLHGSWES--FTGLHTGLD--ASDADADKTLNVMFAANYWHQKGAPR : 261
LOC105328391 chi3l1-2 : GGWSATSGPYSTMASSALTRS-----VFINSLMAWLRQHNFDGVDMDWEFPAYALRGGIPADYTNFALLIKEMRQAFNAEAQSSGRSRLIISAAVSHAKTVIDAGYNVPQIAMYIDFIFIMAYDMHGPWEG--KTGLHSGLY--ASNNDPDKKLNTAWAANYWNSKGMPK : 257
LOC105332575 pchi : GGWDKGSEGFSKLVSSRENML-----KFTKWVIIYLRRFDFDGLDFDWQFPAL--RDSSLQDKKRFADLCDVIALEFDIEEIPDIKWKLTFTLTFHPLLPVMHFGYDIPRLSSKAHYINLKMYDLYGHWDDPLLARHHSALT------GPKGFNNINDLVEMWEEKGVPG : 258
LOC105332577 pchi1 : GGWMMDSEAWSNVVRDKESMD-----TFTANAITFLRFHDFDGIDLDWQYPAF--RGSSHEDQARLVELCERFRHGIETESEPDSKWHLSFSLSVDPSAHRVMYSYDLKRLAKEVDFFNLKTYDLHGHWSEPLTADHHSPLVG-GDELSPEPKDSVNELAKEWVMSGVPA : 260
LOC105332622 amcase-1 : QKLNVGLALYGRSFTLSD-PYNNGVG---APAKGKGNAGNYTREAGFLSYFEVCQKIS--------------EGAQKHYI-PEQESPYITYAD--Q----WVGYDDADSLAVKV-EYIKKQNYGGIMVWALDLDDFN-LICNATTETYPLLRRINQQLGVVIPHGSSSIS : 406
LOC105331571 chit1l : EKLILGLATYGRSFKLQD-ENNFGVG---APATGAGPQGKYVAEDGFLPYYDICARQV--------------QRVGETYRDEKAQTPFFVQEN--I----WVGFDDQLSIYTKVNDLVISKQLGGAMIWALDFDDFN-NICGYGK--YPISRVMTDTLLASESDISITPP : 398
LOC105347316 ctbs-1 : DKLLIGIPWHGYDYTCVNYTTDVKTG-------GKNQMTCYFDERNNTESCNTSLREKMSLAEIKNSYPDQYKNAKSHHYHPIYRSDYINVKKGEQQHQIW--YEGHDSLLDKY-MYVRSMNLRGMVIWTAD--DLS-TKSTDDAAEWNWI------LHNLFSTGDIE-- : 387
LOC105348542 ctbs-2 : NKLVLGVPWYGYIYQCVSITKDNKCAIKHVPFRGVNCSDAAGHEINYS-YIMASLLPK--------------SSTGRIW-DRETLSPRFNFVNNTITYQIW--YDDPESLALKY-NISTYHGLRGVGMWNADAVDFA-----------------------------IDPQ : 357
LOC105331570 amcase-2 : DKLNIGIATYGRSYTLADPQTDRHIG---ASAVAPGRPGQYTRETGFLSYYEICQMQD--------------RRNGKIFRDEEQSVPYYVTGN--T----WVGYDDIYSVKNKA-KWILQEGYGGSMVWSLSLDDFN-KMCSTSSRTYPLTSLISETITNNRSGEMLTTT : 461
LOC105338914 chi3l1-1 : EKLVMGIHGAGSSFTLAN-NSNHGIG---DAVSGGGEKGPLLKFAGRLAYPEICKMRK--------------E---ETVYDEEQKQKYTYYGS--Q----WVGYEDPDTILMKV-QYAINNNFSGVMFWSLDLDDFLGSHCHQGK--YPLLSVVKKVSELLAQNTTPDPA : 440
LOC105331572 chit1 : NKLIIGLGTYGRSFTLVD-TSNHGVG---APASGAGAPGTYTREKGFLSYYEICEMRK--------------NG-GTTYRDPVAKVPYYVNPNTKL----WVGYDDKESIYAKLTELIIKDGFGGAMTWALDLDDFTGTICGEGK--YPLISLMKNVLDQANGGGTVKPP : 406
LOC105328391 chi3l1-2 : SKIVVGIPTYGKSFILDN-AVNNGVG---APTTGMGPAGRFTGEAGALSYYEICEMQQ--------------RGSGTTRRDLAAKVPYFVNTNTKL----WVGFEDKQSVRDKIRELVVGQGYAGAMTWALDLDDFSGSFCAQGS--YPLISTMKVELGSGGGSSSSTSS : 403
LOC105332575 pchi : HKMVLGMPFFGKSYRLED-PAISMPG---APAIGPG-----TDNGDGIPLKHLCHLIR--------------SGLNESFI-PSQQVPYLVQGD--E----WIGYDNQRSIMNKA-TFVFNKPLAGVLLYSLDMDDHK-GACGAGK--FPMSMSVIHGLQAYMLYMQRKHM : 394
LOC105332577 pchi1 : SQIVMGLAFYGRSFKLKS-ANYSFPG---APAVGPGSDG-----GEGIPVNKICHLIR--------------GGVEEKYI-PEKRVPYYVIDD--E----WVGFDNPRSIREKA-QLVAKNHLGGIAMWTMDQDDHR-GVCGEK---WPLMFAIIHGLGPLEYVSYVSAK : 395
LOC105332622 amcase-1 : RAPVSKTQPEVVKIQTSQRASQSDLLRTN-----------------------------------------------------------KPKMSQTSKPIWFKRNQETVRTATVTRPPSGHSSGSNSGSHSTQNSNKPSGGKEF---------KCEVGGDGYYPSPYSCEE : 508
LOC105331571 chit1l : -------------------------------------------------------------------------------------------------------------------------------------------------------------------------- : -
LOC105347316 ctbs-1 : -------------------------------------------------------------------------------------------------------------------------------------------------------------------------- : -
LOC105348542 ctbs-2 : -------------------------------------------------------------------------------------------------------------------------------------------------------------------------- : -
LOC105331570 amcase-2 : TTTTTPSSTTTSTSTTVPTTTTEAETTTS-----------------------------------------------------------MKTTTTTTTPTPLPTTTTTTATTTTTARRIPPTPPPLPPIPEIRTTVKEPSTLPP-----------------IVPVDNGATG : 555
LOC105338914 chi3l1-1 : -------------------------------------------------------------------------------------------------------------------------------------------------------------------------- : -
LOC105331572 chit1 : TTKEPGTNPPTNAPTNAPTTTRAPSVTTPGGGGADGICAQSYHGYIFPNVNDCSSFYQCVHGRAVVIQCQSGLLFNVATDNCDWPSNVVCASTTQAPVTAVPTTQAPVTAVPTTQAPVTAVPTTQAPVTAVPTTIVPSSSKPSSMVPTTPTPAGTCPYDGAIRDPTDCAK : 576
LOC105328391 chi3l1-2 : -------------------------------------------------------------------------------------------------------------------------------------------------------------------------- : -
LOC105332575 pchi : -------------------------------------------------------------------------------------------------------------------------------------------------------------------------- : -
LOC105332577 pchi1 : -------------------------------------------------------------------------------------------------------------------------------------------------------------------------- : -
LOC105332622 amcase-1 : YYMCNDG--HAHHFKCAAGLLFNSQFNYCGWPKDVVCDLDAARKKAKEALKKPESQQQIPQQPVQQNPPPVYKTPAENQNAFQFKPQKPQQPVQQTQQVQPQQVQQQPQTRKQPNTAPQPPQNNVRPPQQASQPNNAMG----SAASNDPWNWSPFQQQQQNPVGLPFYW : 672
LOC105331571 chit1l : ------------------------------------------------------------------------------------------------------------STHIPLSTVGTTTRSPYPPPTGDGGKTPDSG--GGGSGHDGITSLD---------------- : 442
LOC105347316 ctbs-1 : --------------------------------------------------------------------------------------------------------------HHSLDMAGTIAGISVGCFVLGTLLGFAFG------------------------------- : 416
LOC105348542 ctbs-2 : -------------------------------------------------------------------------------------------------------------------------------------------------------------------------- : -
LOC105331570 amcase-2 : YRTLPPSVNVTSSTESTTKVVNKVITVMFINKKELPLAHRTTNKVV---------------------------------------TQTPPTTVTTGEVINFGEVTIAPTPRTTKRPTRTITTTKRPMTTRRLTTAQPIGPFNPNTGRNGFLKNSPLWLVITERKYFPRYS : 686
LOC105338914 chi3l1-1 : --------------------------------------------------------------------------------------------------------------HSQPIKTKPTTTTVWPP------------------------------------------- : 457
LOC105331572 chit1 : YYLCWDGVVLRQPKSCPTGMLFSSVTKLCDRAGVVNCNSDPSSTSA---------------------------------------PTTKQPTTIKPTTTISMKTTTKPTTSTTTTTTKQTTTTPKPPTTKPTTAVPATG--RPTSAP------G---------------- : 683
LOC105328391 chi3l1-2 : --------------------------------------------------------------------------------------------------------------TTTPTTITTTSGGAVS-------------------------------------------- : 419
LOC105332575 pchi : --------------------------------------------------------------------------------------------------------AIGEIYRTKINSARVSRRAYFMAGKKDMVAKEDKKIAG---------------------------- : 432
LOC105332577 pchi1 : ------------------------------------------------------------------------------------------------------HESLRELVNKKIIHANAQFAYYSEAFDSRNARKWEQK------------------------------- : 432
LOC105332622 amcase-1 : SFFRTDLSQDFCTGRANGLYPDPDNCQGFIDCVREIS-FKGLCSPGMAFDTARHMCDY-------------------------------IQNVPGCNDASPM--------------------- : 742
LOC105331571 chit1l : --VD-------CGHEGDGLYRYLSDCSKYIQCVKGKT-FVRNCPTDLEFNIAFSQCDWASNVNCSSILVTIPKTTTSVYSKTDNSSLNITPINTGCQLSPYSHHLTLYTYMYLLIFFLAISMS : 555
LOC105347316 ctbs-1 : -----------CAVMRRRMQRKG----------LRLP-FHRDEEETDFHDDDNTL-------------------------------------------------------------------- : 449
LOC105348542 ctbs-2 : --------------GAKGMW-------------DALPVYPRNSS------------------------------------------------------------------------------- : 374
LOC105331570 amcase-2 : RYFS-------CHRRRNGYYVDVHDCSKYYVCQNRRT-FHYKCPKDLYYNPRKQICDW--------------------------------ISQVSCVNLPR---------------------- : 747
LOC105338914 chi3l1-1 : --------------KDGGFKLFI------IDSRNSTPLLTFPC---LILLLLFSLCVL-------------------------------FINHP----------------------------- : 497
LOC105331572 chit1 : --IS-------CTNNEGKLYPYSKDCTKYVQCVHGSP-LVRPCNPGLHFSPTSLTCTW--------------------------------SNLAGCQ-------------------------- : 738
LOC105328391 chi3l1-2 : -----------CVGNEGALLPHP-DCTKYYQCVNGIP-MEKSCPGGLHFSVTISACDW--------------------------------PSNAGCQ-------------------------- : 471
LOC105332575 pchi : -----------LQKEQDIAMGKMTAEVQSAQAMMNKP-FELPSPNQRSFN-------W----------------------------------------------------------------- : 471
LOC105332577 pchi1 : -----------LAVLTTKLADMQAQQSVFWSKVQGAA-TRGGSPMPPIDKPSWTM-------------------------------------------------------------------- :
图 3 长牡蛎 Cgchitinase 家族蛋白氨基酸多序列比对
图中红框所示为 Glyco-18 结构域,黑色区域为高度相似区域,深灰色为较高相似区域,灰色为相似区域。
Fig. 3 Multiple sequences alignment of the Cgchitinase proteins in C. gigas
The red box in the figure showed the Glyco-18 domain, the black area was a highly similar area, dark gray was the higher similarity area, and gray was
the similar area.
而 长 牡 蛎 的 amcase-1 与 uncharacterized protein 及 chitinase10聚为一支并最终形成一大
LOC105332578 分 别 与 红 鲍 的 uncharacterized 分支 (图 4)。
https://www.china-fishery.cn 中国水产学会主办 sponsored by China Society of Fisheries
6